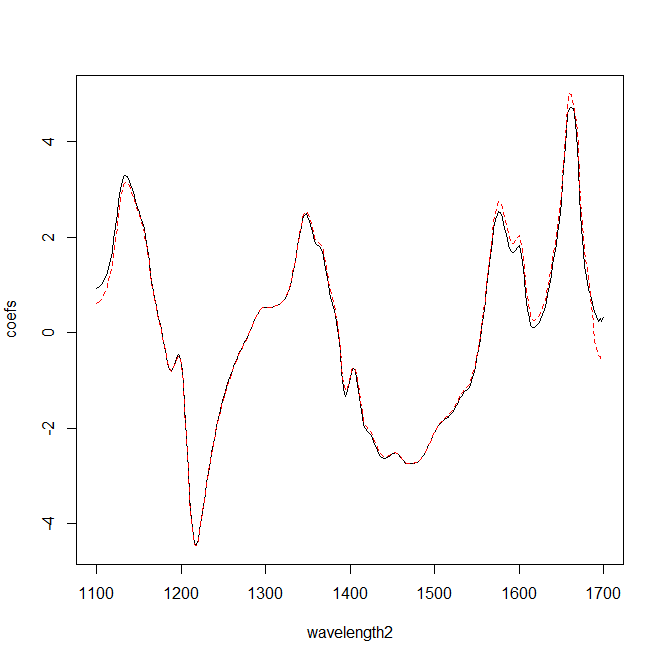
I was practicing with the shootout 2002, where you have a certain number of training samples (155), scanned in two instruments 1 and 2, so we have two "Training files" with the same LAB values, but different spectra (due that the samples are acquired in diferent instruments). The idea of the shootout is to develop a robust calibration for both instruments.
So I had developed a PLS Model in R with the Training samples acquired in "Instrument 1". I called the model "mod1":
mod1<-plsr(Y~X1,data=nir.training1,ncomp=5,validation="LOO").
Where "nir.training1" is a data frame:
nir.training1<-data.frame(X= I(X1),Y=I(Y))
X: is the 155 row training matrix of spectra acquired in Instrument 1
Y: is the reference training matrix of the constituent of interest and this matrix is the same for the Instrument 2, where their matrix of spectra would be X2.
after checking the summary of the model I decide that 3 terms (components) are enough for the model.
Now I want to predict the samples of a Test file (also scanned in 1). This Test file has more samples (460), so the rows are higher for the Y matrix and for the X matrix. The idea is to get a RMSEP statistic.
RMSEP(mod1,estimate="test",ncomp=3,intercept=FALSE,
+ newdata=data.frame(Y=I(Y.test),X=I(X1.test)))
and I have found several problems getting errors like this:
"newdata' had 460 rows but variables found have 155 rows"
"Error en model.frame.default(formula(object), data = newdata) :
variable lengths differ (found for 'X1')".
I was making some mistakes assigning names to the matrix in the data frame, and they must be the same in both cases (The dataframe from which I develop the regression and the dataframe which I want to evaluate.
nir.training1<-data.frame(X= I(X1),Y=I(Y))
newdata=data.frame(Y=I(Y.test),X=I(X1.test)))
Finally I got a value of 4.974 for the RMSEP.
Now the following exercise must be to check if I have a similar error with a calibration develop with a model developed with the Training spectra of Instrument 2 with the Test Set spectra from Instrument 2 (don´t forget that the Y reference values are the same that for instrument 1, because we are using the same samples):
mod2<-plsr(Y~X,data=nir.training2,ncomp=5,validation="LOO")
Three terms are also enough, and the RMSEP for the Test Set scanned in Instrument 2 with the model 2 is:
RMSEP(mod2,estimate="test",ncomp=3,intercept=FALSE,
+ newdata=data.frame(Y=I(Y.test),X=I(X2.test)))
and the value is: 5,315.
But what would be the RMSEP for the Test set scanned in Instrument 2 and predicted with the model developed with the Training Samples scanned in instriment 1 (mod1):
RMSEP(mod1,estimate="test",ncomp=3,intercept=FALSE,
+ newdata=data.frame(Y=I(Y.test),X=I(X2.test)))
The result is: 9,983
So the model can not be transfered without doing anything from instrument 1 to Instrument 2, if what we want is to get similar performance in both.
We will continue with this in the next post.