Following a course about Machine Learning with R, I realize of the importance of the random selection of the samples for the development of the models.
R has good tools to select the samples randomly and to do it was a common practice during all the exercises in the course.
I do it, in the case with the soy meal samples I have used for several posts, so we will compare the results.
The idea of random selection is to make the model robust against the bias which we see quite often when validating the models with independent samples.
We can see if the number of the terms selected change or if we get similar results to the previous selection using an structure selection of odd and even samples.
Random selection order is important also for a better cross validation.
Here is the code and preparation of the sample sets for the development of the models.
##################### SPLITTING SAMPLES RANDOMLY #############
#In this case we need first the dataframe "soy_ift_conv"
#Split the data into 65% training and 35% test
rndIndices=sample(nrow(soy_ift_conv))
sepPnt=round(.65*nrow(soy_ift_conv))
train=soy_ift_conv[rndIndices[1:sepPnt],]
validation=soy_ift_conv[rndIndices[(sepPnt+1):length(rndIndices)],]
#Plotting Training and Validation sets overlapped.
matplot(wavelengths,t(train$X_msc),type="l",
xlab="wavelengths",ylab="Absorbance"
,col="blue")
par(new=TRUE)
matplot(wavelengths,t(validation$X_msc),lty=1,
pch=NULL,axes=FALSE,
type="l",col="gray",xlab="",ylab="")
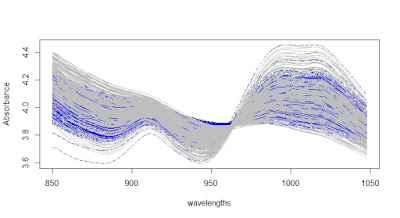
We see in gray the validation samples selected and in blue the training samples.
No hay comentarios:
Publicar un comentario